|
20 Here, we present a de novo assembly approach for transcriptome of a plant species using only short-read sequence data. Recently, the de novo assembly of human transcriptome has been reported using ABySS program. 15, 16 Although the efforts have been made to develop tools for de novo assembly using short-read sequence data, 17–19 their use in transcriptome assembly has not been well demonstrated yet. However, most of these studies are based on the long-read sequence data using 454 pyrosequencing or employing hybrid approach. 14 Several studies have reported the transcriptome sequencing of various model and non-model species using these technologies. The next generation sequencing technologies provide a cost-effective means of sequencing the transcriptome of an organism. This is reflected by a very small number of ESTs (34 587) present in the dbEST database at NCBI (release 100110 1 October 2010) for chickpea. Although ESTs and other cDNA sequences are among the most reliable evidences for the identification of gene-rich regions in a genome, gene identification and genome annotation, very less effort has been made for chickpea in this direction when compared with other crop plants. The generation of large-scale ESTs is a very useful approach to accelerate the research on non-model species. 2 The function of a few genes in stress responses has also been demonstrated using transgenic approach. 10, 11 The efforts have been made to clone genes of interest via candidate gene approach. 5–9 In addition, other microarray and SAGE technologies have also been used to identify the stress-responsive transcriptome in chickpea. Most of the ESTs have been generated with the aim to identify the candidate genes involved in various abiotic and biotic stress responses and development of molecular markers. 3, 4 Although a few expressed sequence tags (ESTs) have been generated and gene-based markers have been developed, the functional genomics studies in chickpea is still in its infancy. Various genomic tools have facilitated greatly the development of improved genotypes/varieties in several crop species. 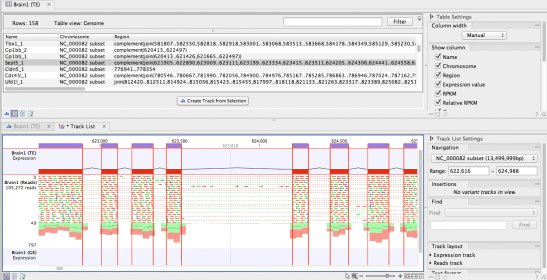
2 Unfortunately, very limited genomic information is available for chickpea. Modern breeding technologies with biotechnological techniques are required to increase the productivity. Several biotic such as Ascochyta blight, dry root rot, Fusarium wilt and pod borer, and abiotic such as drought, salinity and low temperature, constraints are major factors for lower chickpea production. Despite growing demand and high-yield potential, chickpea productivity is very low. 1 Chickpea is a self-pollinated, diploid (2 n = 2 x = 16) and annual plant with a moderate genome size of about 740 Mb. IntroductionĬhickpea ( Cicer arietinum L.) is the third most consumed legume crop, which is cultivated in arid and semi-arid areas around the world. In addition, the strategy for de novo assembly of transcriptome data presented here will be helpful in other similar transcriptome studies.ĭe novo assembly, chickpea, next generation sequencing, transcriptome, short read 1. The chickpea transcripts set generated here provides a resource for gene discovery and development of functional molecular markers. 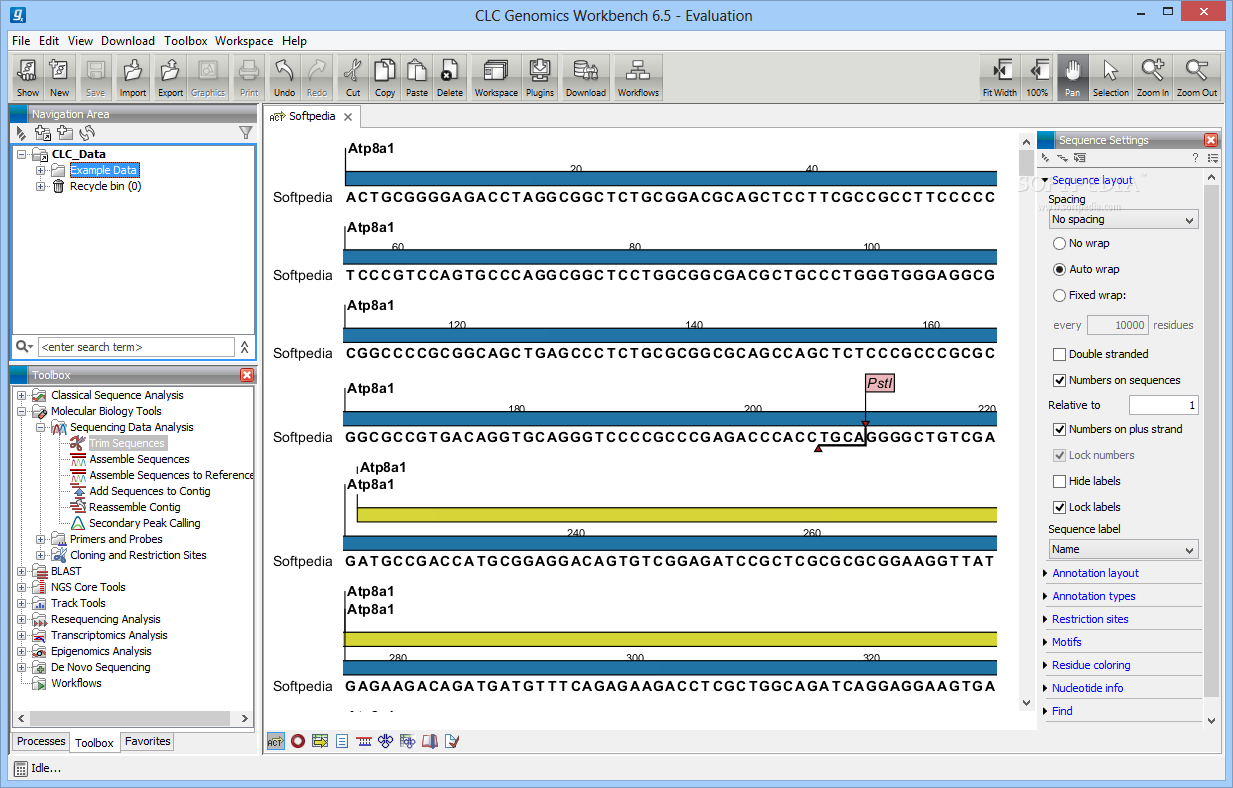
In addition, we identified simple sequence repeat motifs in transcripts. Functional categorization revealed the conservation of genes involved in various biological processes in chickpea. At the protein level, a total of 45 636 (85.5%) chickpea transcripts showed significant similarity with unigenes/predicted proteins from other legumes or sequenced plant genomes. The average length of transcripts was 523 bp and N50 length of 900 bp with coverage of 25.7 rpkm (reads per kilobase per million). We assembled ∼107million high-quality trimmed reads using Velvet followed by Oases with optimal parameters into a non-redundant set of 53 409 transcripts (≥100 bp), representing about 28 Mb of unique transcriptome sequence. We have assessed the effect of sequence quality, various assembly parameters and assembly programs on the final assembly output. 
In the present study, the transcriptome of chickpea was sequenced with short reads on Illumina Genome Analyzer platform. However, the genomic resources available for chickpea are still very limited. Analyzing and visualizing Next Generation Sequencing data, incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of your typical NGS workflow.Chickpea ranks third among the food legume crops production in the world.The CLC Genomics Workbench is the client software for the CLC Genomics Server.įeatures for the Genomics Workbench are described below:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
